
I visited the one and only microbe museum-, Micropia in Amsterdam in August, following the ESEB meeting in Groningen. Highly recommend for anyone with an interest in microbes or science in general. Very interactive.

I visited the one and only microbe museum-, Micropia in Amsterdam in August, following the ESEB meeting in Groningen. Highly recommend for anyone with an interest in microbes or science in general. Very interactive.
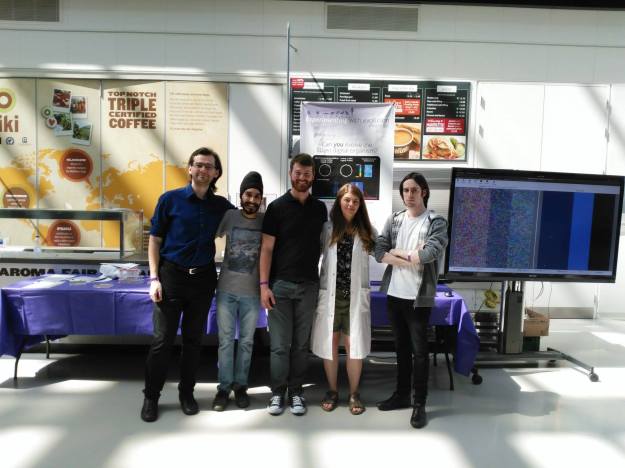

Knight lab does the University of Manchester Community Festival! Live-action demos of evolution simulations and examples of antibiotic resistant E. coli.
This somewhat-ridiculous BASH one-liner will create a BibTeX database file (.bib) from a bunch of PDFs via the Crossref API for DOIs, providing the PDF has a DOI on the first page. As DOI was introduced in 2000, this will probably not work on vintage PDFs.
for pdfs in *.pdf; do pdftotext -f 1 -l 1 "$pdfs" - |tr -d "\n" | grep -oE "(doi|DOI):\s?[A-Za-z0-9./-\(\)-]+[0-9]" | tr '[:upper:]' '[:lower:]' | sed -r 's;doi:\s?;http://api.crossref.org/works/;g' | sed -r 's;$;/transform/application/x-bibtex;g' | xargs curl -fsS 2>/dev/null | sed -e '$a\'; done > allpdf.bib

Setting up a large selection experiment on antibiotic fitness landscapes. I’ve decided to use the Singer ROTOR HDA for consistency and repeatability. The Singer ROTOR HDA uses a pre-sterilized pad system (called RePads) to transfer bacteria between solid or liquid medium. RePads are available in 96, 384, 1536 or 6144 formats. Read more about the Singer ROTOR HDA.
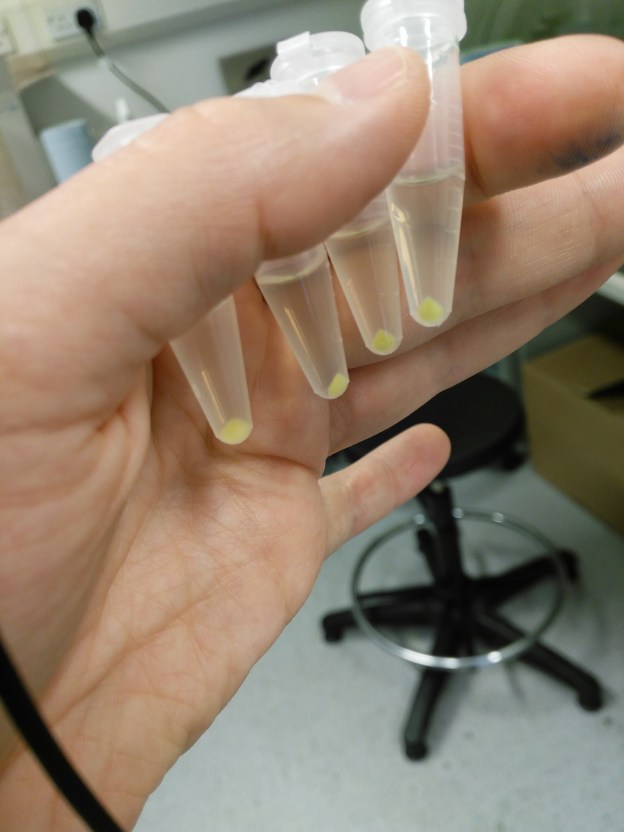
Pelletted E. coli S17-1 with the plasmid pGRG25-Pmax::GFP. they are really green!
Vented Petri dishes have a small lip on the top edge of the dish that allows the lid to sit a little up from the bottom, allowing for some air flow. Non-vented Petri dishes allow the lid to sit more or less flat on the bottom. I was wondering what the best applications are for triple, single and non-vented Petri dishes, and found this guide from Thomas Scientific (link is dead).
Edit 28-02-2019 there’s an even better summary from Tritech Research:
Some of our dish models are available in both “vented” and “non-vented” styles. Standard Petri Dishes are always vented, so if the don’t say vented or non-vented, you should assume they are vented. “Vented” means that the lid is slightly elevated above the base. This allows for good, plentiful air exchange. This is useful when you want to encourage evaporation, for example, when you want to use poured plates as soon as possible, and the plates themselves, or a liquid seeding solution, needs to dry beforehand. The basic design of the dish tends to maintain sterility because particles would have to go up and over the dish’s wall to get inside, and this is rare in normal airflow.
With “non-vented” dishes, the lid fits quite flatly on the base. While it is not a hermetic seal, the space between dish and lid is extremely small. This results in even less potential for external contamination and a significantly reduced evaporation rate. For example a 60mm vented Petri Dish containing 10ml of agar medium typically dries out in 2-3 weeks; whereas, a similar 60mm non-vented dish typically lasts 2-3 months. Most C. elegans labs, except those in very humid climates, prefer the non-vented dishes. Non-vented dishes provide sufficient air exchange for the worms to breathe while greatly increasing the life of the dish.
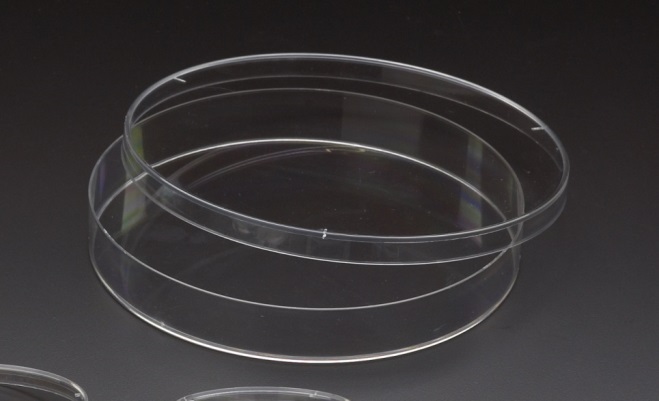

Non-vented Petri dish
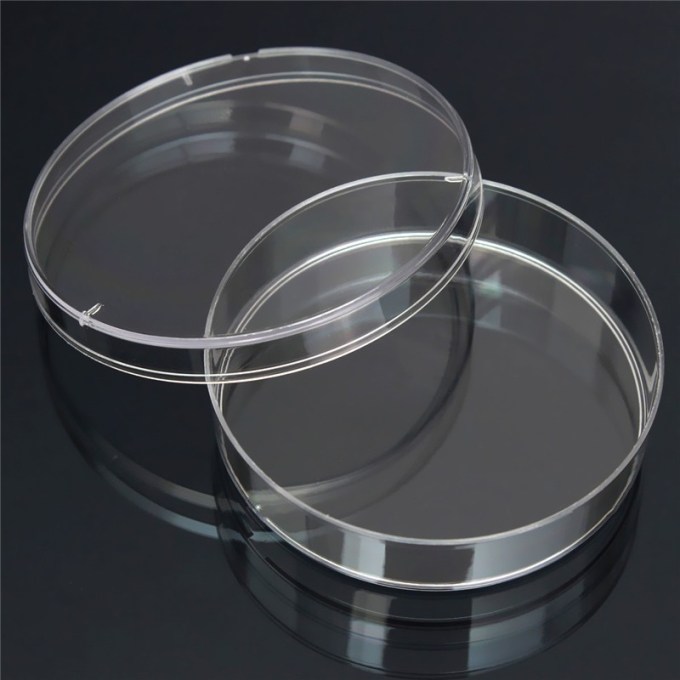

Vented Petri dish
Here’s a quick and dirty Perl script to get a data table out of a .jrp file from JMP. Not guaranteed to work for all files, as I’ve only tested it on one (so modification may be necessary).
#! /usr/bin/perl -w
use strict;
use Getopt::Long;
my $jmp_file;
my $colhead;
my $values;
my $row;
my @records;
my $ndata;
GetOptions ('jmp=s' => \$jmp_file) or die("Error in arguments\n");
open (JMP,"<$jmp_file") || die "cannot open JMP input file $jmp_file";
while(<JMP>){
chomp;
if($_=~/New Column\(\s\"(.+?)\",.+$/){ # Get column names
$colhead=$1;
}
if($_=~/Set Values\(\s[\[\{](.+?)[\]\}] \) \),/){ # Get row values
$values=$1;
my @row = split ", ", $values;
unshift @row, $colhead;
$ndata= scalar @row;
push @records, \@row;
}
}
# Rotate table 90 degrees (rows-to-columns)
my $nrecords=scalar @records;
for(my $i=0;$i<$ndata; $i++){
for(my $j=0;$j<$nrecords;$j++){
print "$records[$j][$i]";
print "," if $j<($nrecords-1)
}
print "\n";
}